Overview
This vignette presents analyst-oriented tools added to betaregscale for post-estimation workflows:
-
brs_table(): model comparison tables. -
brs_marginaleffects(): average marginal effects (AME). -
brs_predict_scoreprob(): probabilities on the original score scale. -
autoplot.brs():ggplot2diagnostics. -
brs_cv(): repeated k-fold cross-validation.
The examples use bounded scale data under the interval-censored beta regression framework implemented in the package.
Mathematical foundations
Complete likelihood by censoring type
For each observation , let indicate exact, left-censored, right-censored, or interval-censored status. With beta CDF , beta density , and interval endpoints , the contribution is:
This is the basis for fitting, prediction, and validation metrics.
Model-comparison metrics
brs_table() reports:
- ,
- ,
- ,
where is the number of estimated parameters and is the sample size.
Average marginal effects (AME)
brs_marginaleffects() computes AME by finite
differences:
with
on the requested prediction scale (response or
link). For binary covariates
,
it uses the discrete contrast
.
Score-scale probabilities
For integer scores
,
brs_predict_scoreprob() computes:
These probabilities are directly aligned with interval geometry on the original instrument scale.
Cross-validation log score
In brs_cv(), fold-level predictive quality includes:
where is the predictive contribution from the same censoring-rule piecewise definition shown above.
It also reports:
Reproducible workflow
1) Simulate data and fit candidate models
set.seed(2026)
n <- 220
d <- data.frame(
x1 = rnorm(n),
x2 = rnorm(n),
z1 = rnorm(n)
)
sim <- brs_sim(
formula = ~ x1 + x2 | z1,
data = d,
beta = c(0.20, -0.45, 0.25),
zeta = c(0.15, -0.30),
ncuts = 100,
repar = 2
)
fit_fixed <- brs(y ~ x1 + x2, data = sim, repar = 2)
fit_var <- brs(y ~ x1 + x2 | z1, data = sim, repar = 2)2) Compare models in one table
tab <- brs_table(
fixed = fit_fixed,
variable = fit_var,
sort_by = "AIC"
)
kbl10(tab)| model | nobs | npar | logLik | AIC | BIC | pseudo_r2 | exact | left | right | interval | prop_exact | prop_left | prop_right | prop_interval |
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| variable | 220 | 5 | -931.8646 | 1873.729 | 1890.697 | 0.1381 | 0 | 8 | 22 | 190 | 0 | 0.0364 | 0.1 | 0.8636 |
| fixed | 220 | 4 | -939.3243 | 1886.649 | 1900.223 | 0.1381 | 0 | 8 | 22 | 190 | 0 | 0.0364 | 0.1 | 0.8636 |
3) Estimate average marginal effects
set.seed(2026) # For marginal effects simulation
me_mean <- brs_marginaleffects(
fit_var,
model = "mean",
type = "response",
interval = TRUE,
n_sim = 120
)
kbl10(me_mean)| variable | ame | std.error | ci.lower | ci.upper | model | type | n |
|---|---|---|---|---|---|---|---|
| x1 | -0.1035 | 0.0178 | -0.1339 | -0.0675 | mean | response | 220 |
| x2 | 0.0734 | 0.0188 | 0.0360 | 0.1071 | mean | response | 220 |
set.seed(2026) # Reset seed for reproducibility
me_precision <- brs_marginaleffects(
fit_var,
model = "precision",
type = "link",
interval = TRUE,
n_sim = 120
)
kbl10(me_precision)| variable | ame | std.error | ci.lower | ci.upper | model | type | n |
|---|---|---|---|---|---|---|---|
| z1 | -0.3361 | 0.0835 | -0.4781 | -0.1413 | precision | link | 220 |
4) Predict score probabilities
P <- brs_predict_scoreprob(fit_var, scores = 0:10)
dim(P)
#> [1] 220 11
kbl10(P[1:6, 1:5])| score_0 | score_1 | score_2 | score_3 | score_4 |
|---|---|---|---|---|
| 0.0146 | 0.0178 | 0.0146 | 0.0130 | 0.0121 |
| 0.0215 | 0.0228 | 0.0178 | 0.0155 | 0.0141 |
| 0.0465 | 0.0318 | 0.0216 | 0.0175 | 0.0151 |
| 0.0731 | 0.0371 | 0.0233 | 0.0181 | 0.0152 |
| 0.0076 | 0.0105 | 0.0090 | 0.0083 | 0.0078 |
| 0.0050 | 0.0050 | 0.0039 | 0.0034 | 0.0030 |
kbl10(
data.frame(row_sum = rowSums(P)),
digits = 4
)| row_sum |
|---|
| 0.1344 |
| 0.1614 |
| 0.2001 |
| 0.2313 |
| 0.0856 |
| 0.0354 |
| 0.0295 |
| 0.0994 |
| 0.1024 |
| 0.1312 |
rowSums(P) should be close to 1 (up to numerical
tolerance and score subset selection).
5) ggplot2 diagnostics
autoplot.brs(fit_var, type = "calibration")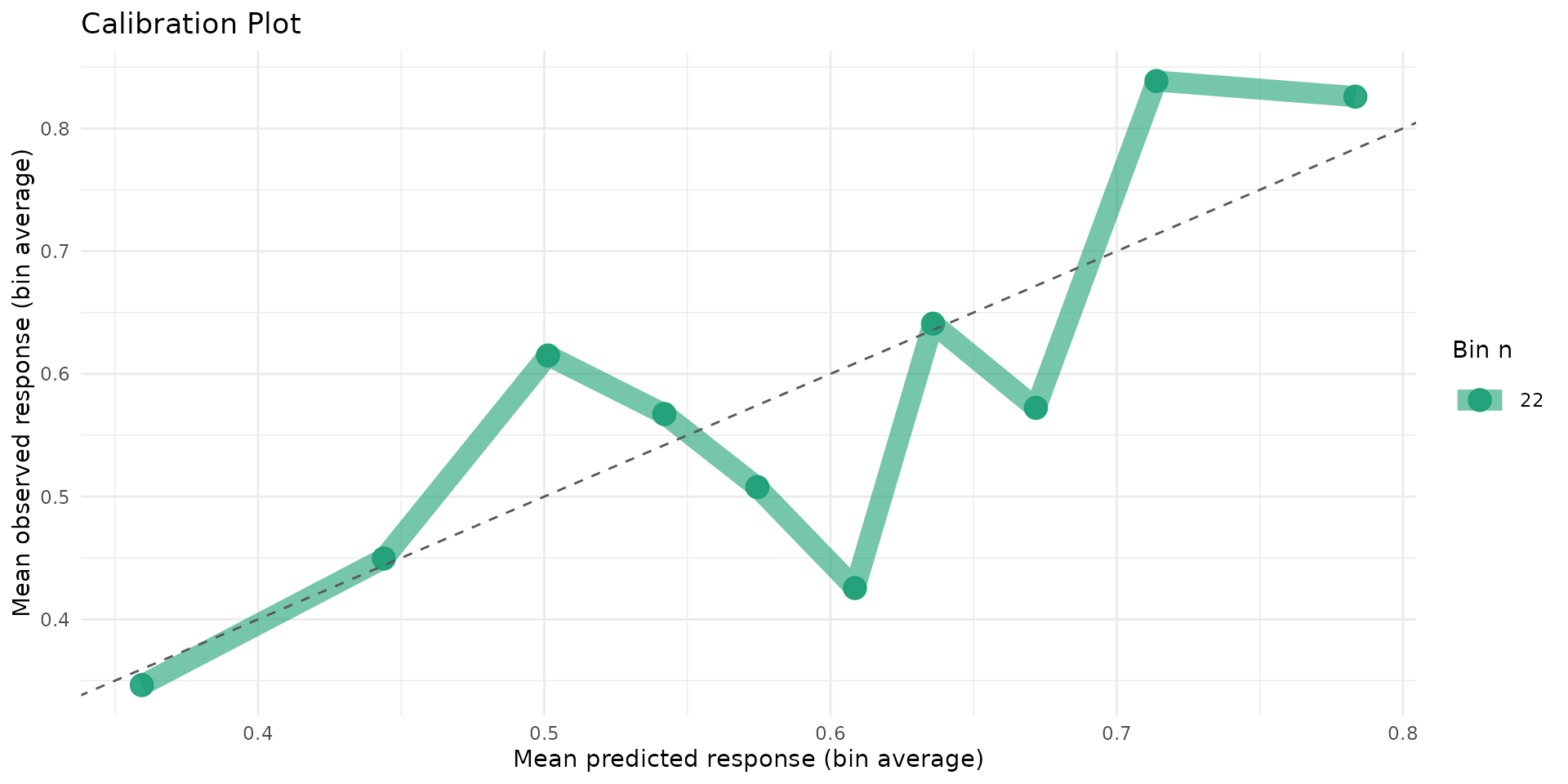
autoplot.brs(fit_var, type = "score_dist", scores = 0:20)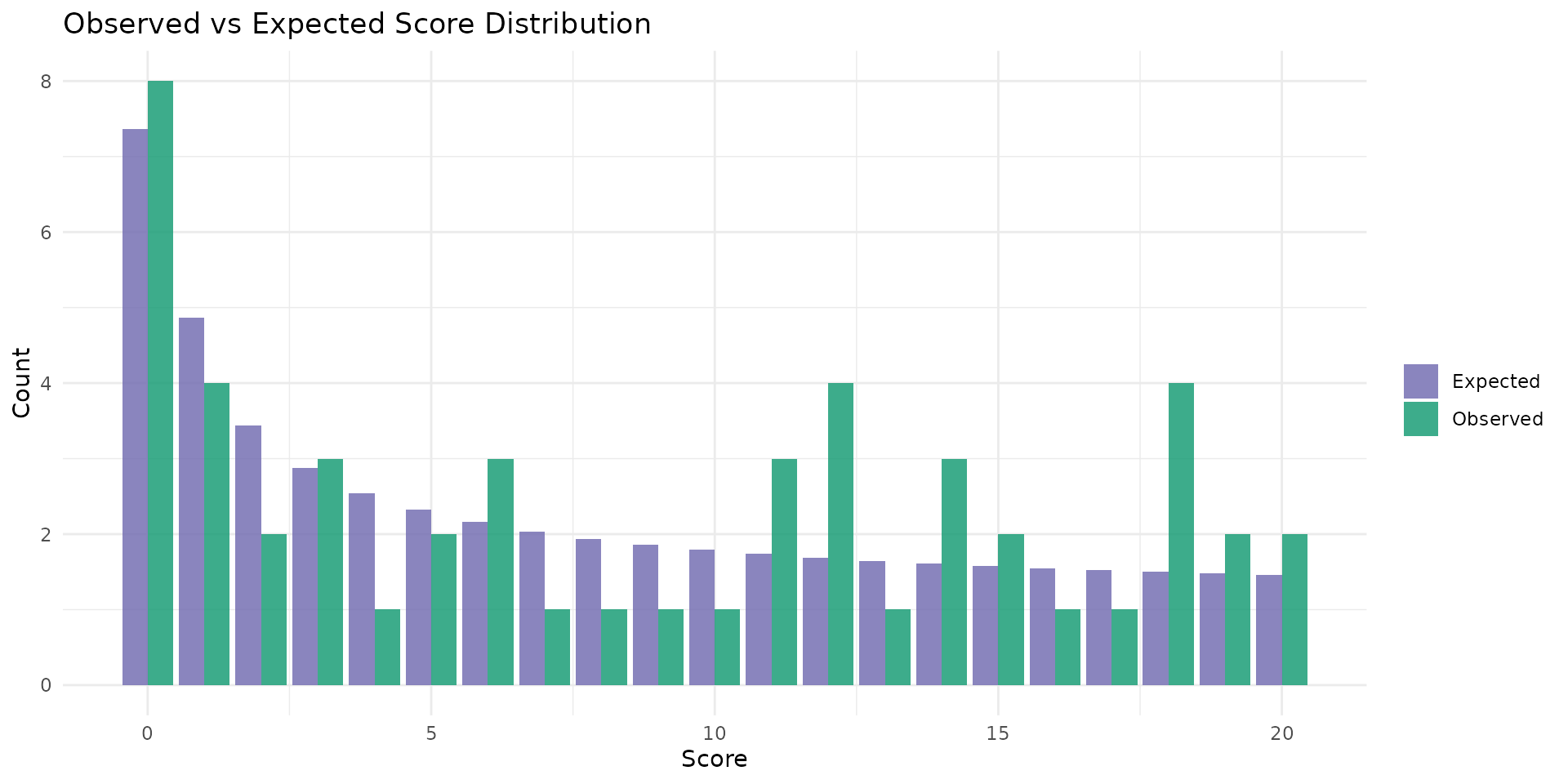
autoplot.brs(fit_var, type = "cdf", max_curves = 4)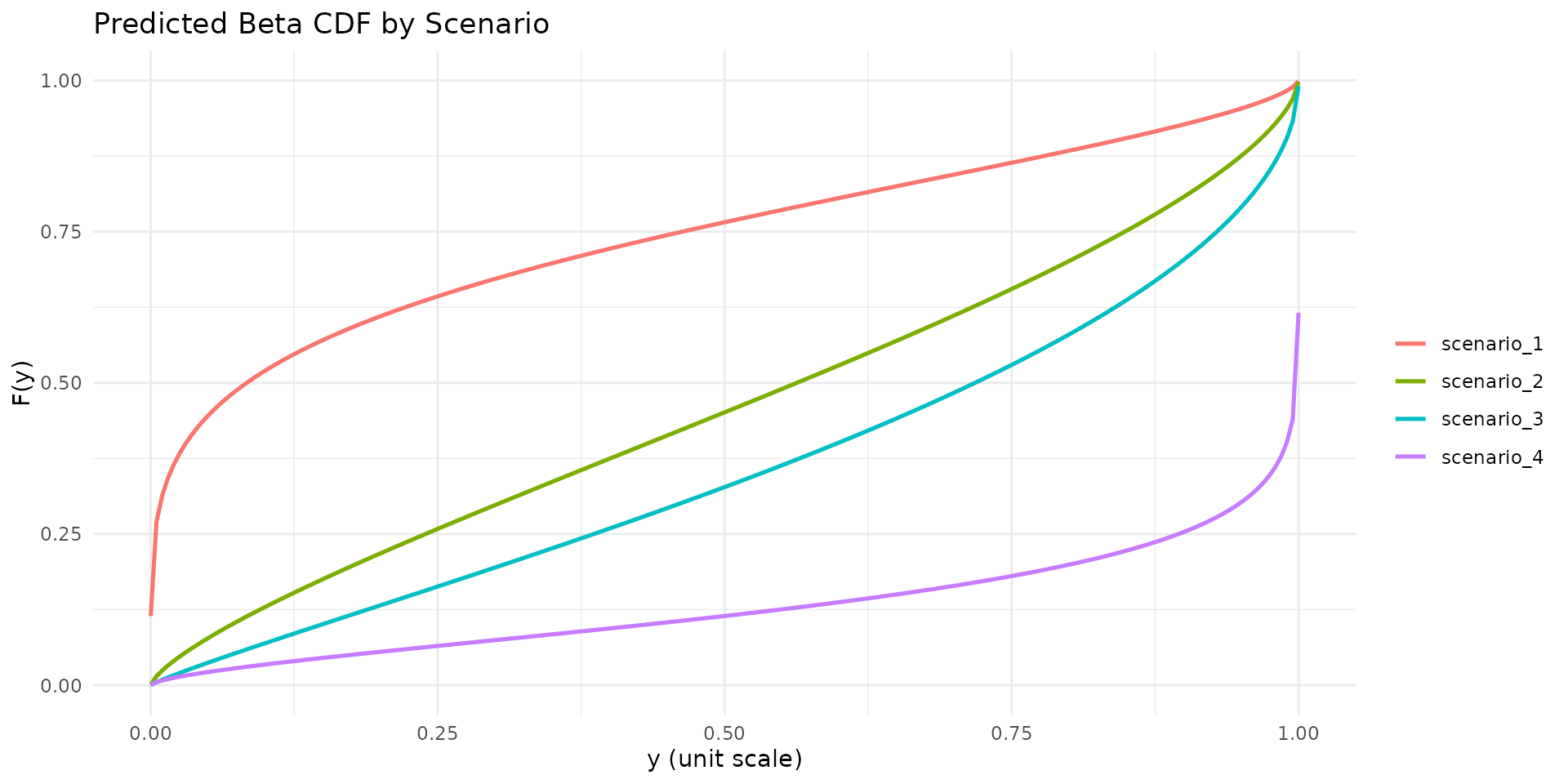
autoplot.brs(fit_var, type = "residuals_by_delta", residual_type = "rqr")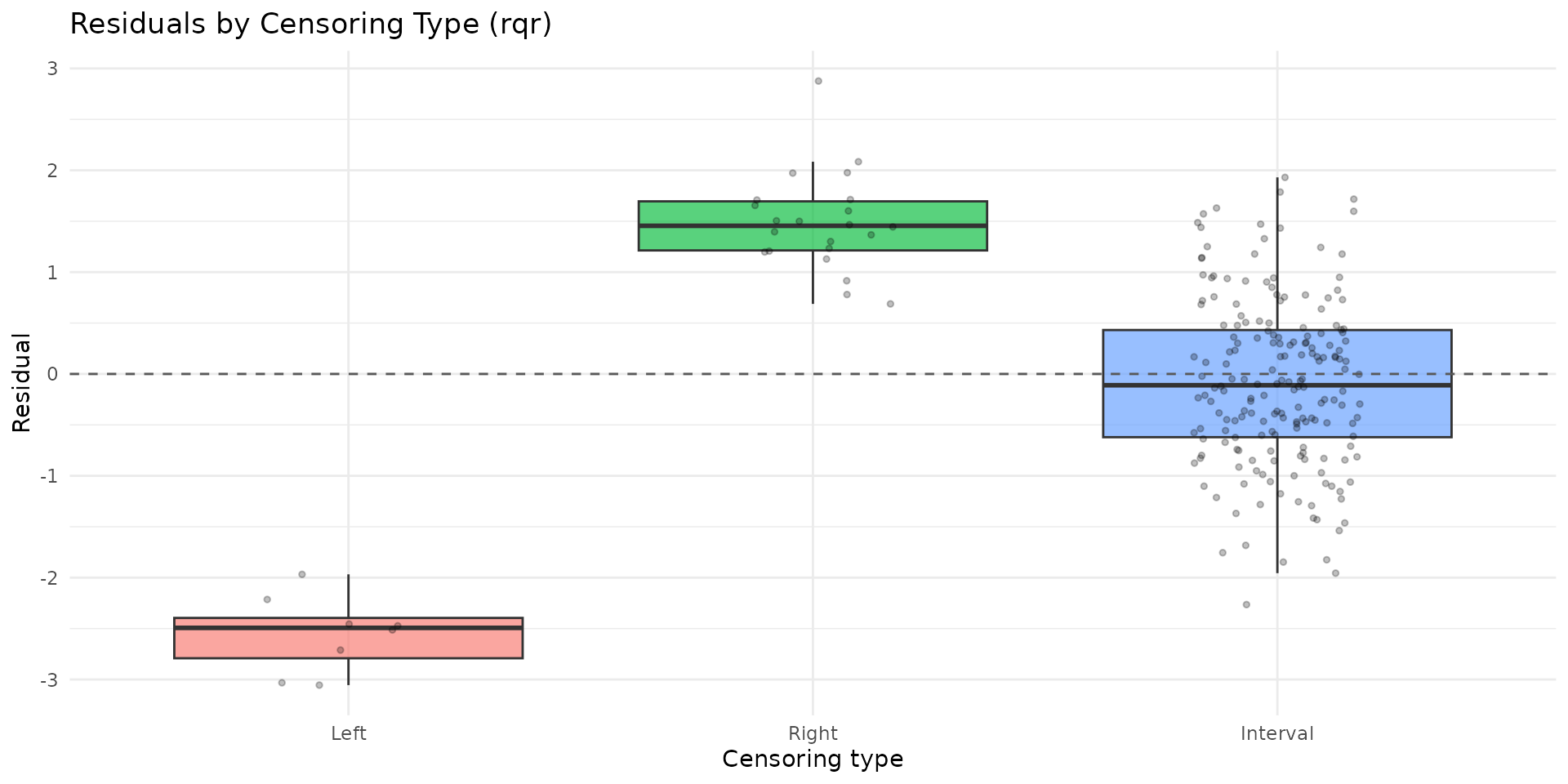
6) Repeated k-fold cross-validation
set.seed(2026) # For cross-validation reproducibility
cv_res <- brs_cv(
y ~ x1 + x2 | z1,
data = sim,
k = 3,
repeats = 1,
repar = 2
)
kbl10(cv_res)| repeat | fold | n_train | n_test | log_score | rmse_yt | mae_yt | converged | error |
|---|---|---|---|---|---|---|---|---|
| 1 | 1 | 146 | 74 | -4.3077 | 0.3215 | 0.2862 | TRUE | NA |
| 1 | 2 | 147 | 73 | -4.3171 | 0.3449 | 0.3032 | TRUE | NA |
| 1 | 3 | 147 | 73 | -4.1676 | 0.3454 | 0.3022 | TRUE | NA |
kbl10(
data.frame(
metric = c("log_score", "rmse_yt", "mae_yt"),
mean = colMeans(cv_res[, c("log_score", "rmse_yt", "mae_yt")], na.rm = TRUE)
),
digits = 4
)| metric | mean | |
|---|---|---|
| log_score | log_score | -4.2641 |
| rmse_yt | rmse_yt | 0.3373 |
| mae_yt | mae_yt | 0.2972 |
Practical interpretation
- Prefer the model with the lower AIC/BIC and better predictive
log_score. - Use AME on the response scale to communicate the expected change in mean score (on the unit interval) resulting from small covariate shifts.
- Use score probabilities to translate model outputs back to clinically interpretable scale categories.
- Inspect calibration and residual-by-censoring plots before proceeding with statistical inference.
References
- Ferrari, S. L. P., and Cribari-Neto, F. (2004). Beta regression for modelling rates and proportions. Journal of Applied Statistics, 31(7), 799-815. DOI: 10.1080/0266476042000214501. Validated online via: https://doi.org/10.1080/0266476042000214501.
- Hawker, G. A., Mian, S., Kendzerska, T., and French, M. (2011). Measures of adult pain: VAS, NRS, MPQ, SF-MPQ, CPGS, SF-36 BPS, and ICOAP. Arthritis Care and Research, 63(S11), S240-S252. DOI: 10.1002/acr.20543. Validated online via: https://doi.org/10.1002/acr.20543.
- Hjermstad, M. J., Fayers, P. M., Haugen, D. F., et al. (2011). Comparing NRS, VRS, and VAS pain scales in adults: a systematic review. Journal of Pain and Symptom Management, 41(6), 1073-1093. DOI: 10.1016/j.jpainsymman.2010.08.016. Validated online via: https://doi.org/10.1016/j.jpainsymman.2010.08.016.