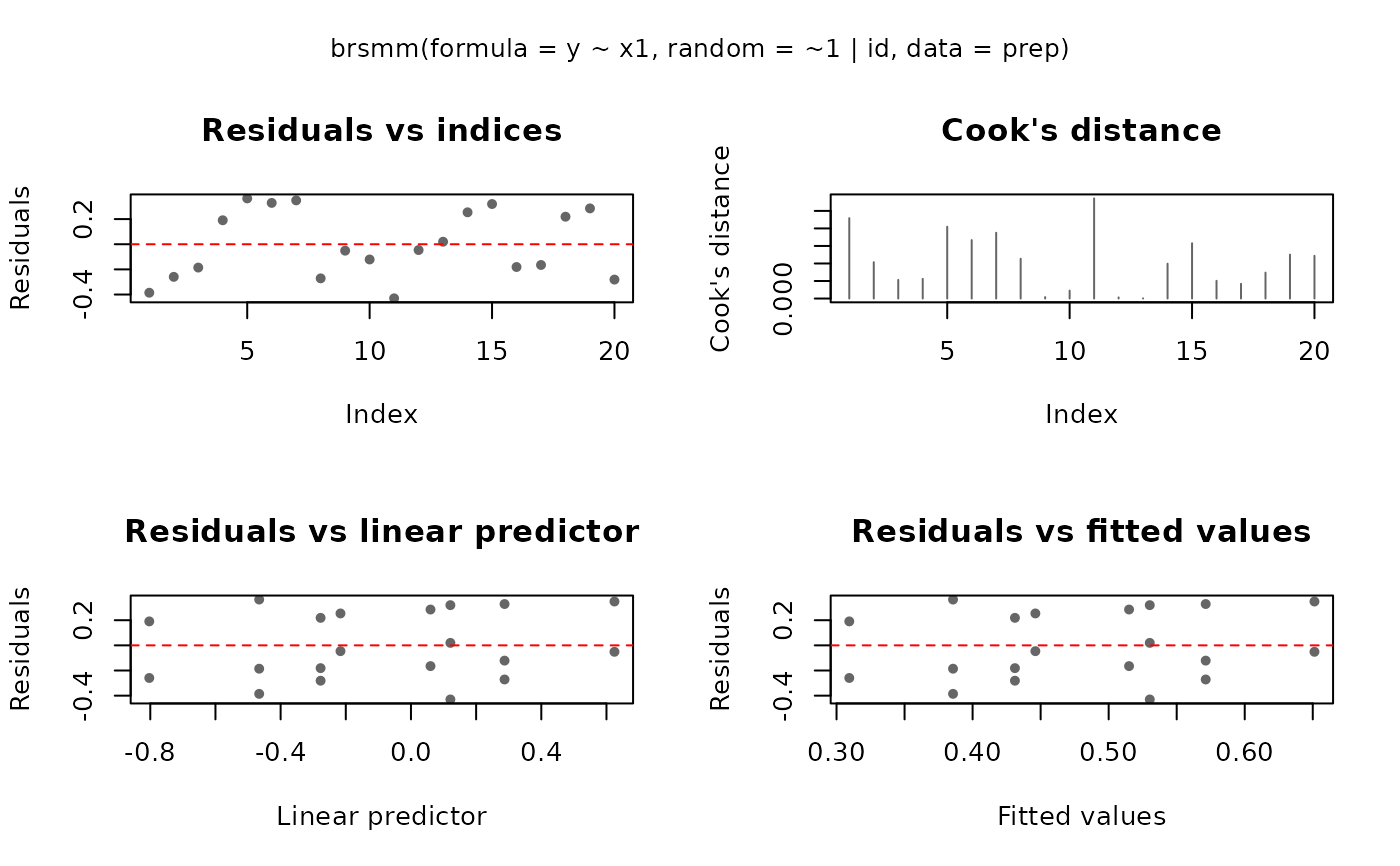
Produces diagnostic plots for fitted "brsmm" models:
residuals vs indices, Cook's distance, residuals vs linear predictor,
residuals vs fitted values, half-normal envelope, and predicted vs observed.
Usage
# S3 method for class 'brsmm'
plot(
x,
which = 1:4,
type = c("response", "pearson"),
nsim = 100L,
level = 0.9,
caption = c("Residuals vs indices", "Cook's distance", "Residuals vs linear predictor",
"Residuals vs fitted values", "Half-normal plot", "Predicted vs observed",
"Random-effects Q-Q", "Random-effects caterpillar"),
sub.caption = NULL,
ask = prod(par("mfcol")) < length(which) && dev.interactive(),
gg = FALSE,
title = NULL,
theme = NULL,
...
)Arguments
- x
A fitted
"brsmm"object.- which
Integer vector selecting which panels to draw (default
1:4).- type
Residual type passed to
residuals.brsmm("response"or"pearson").- nsim
Number of simulations for half-normal envelope.
- level
Confidence level for the half-normal envelope.
- caption
Character vector of plot captions.
- sub.caption
Optional subtitle; defaults to model call.
- ask
Logical: prompt before each new page?
- gg
Logical: use ggplot2 backend?
- title
Optional global title for ggplot output. If
NULL, panel captions are used.- theme
Optional ggplot2 theme object (e.g.,
ggplot2::theme_bw()). IfNULL, a minimal theme is used.- ...
Further arguments passed to base
plot().
Examples
# \donttest{
dat <- data.frame(
y = c(
0, 5, 20, 50, 75, 90, 100, 30, 60, 45,
10, 40, 55, 70, 85, 25, 35, 65, 80, 15
),
x1 = rep(c(1, 2), 10),
id = factor(rep(1:4, each = 5))
)
prep <- brs_prep(dat, ncuts = 100)
#> brs_prep: n = 20 | exact = 0, left = 1, right = 1, interval = 18
fit <- brsmm(y ~ x1, random = ~ 1 | id, data = prep)
plot(fit, which = 1:4)
 # }
# }